|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|

|
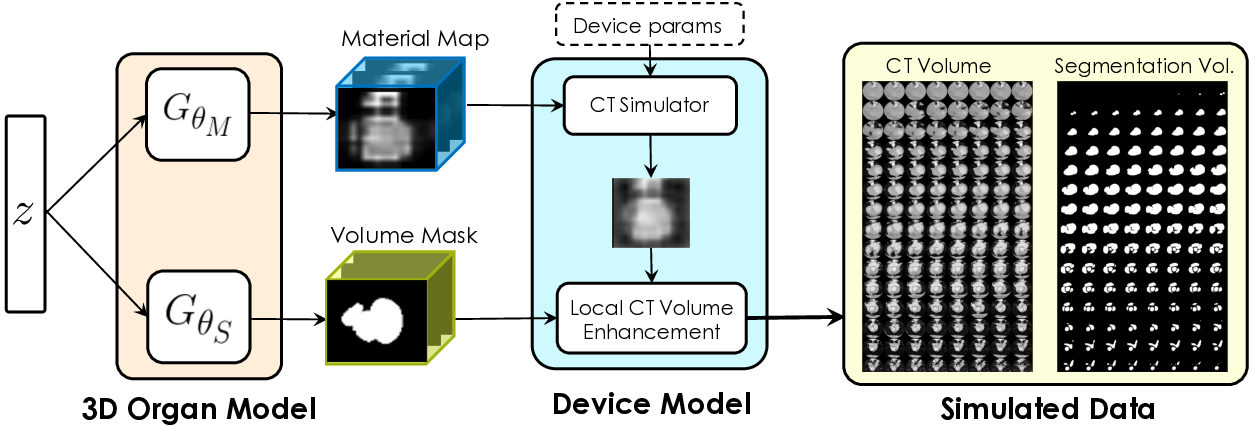
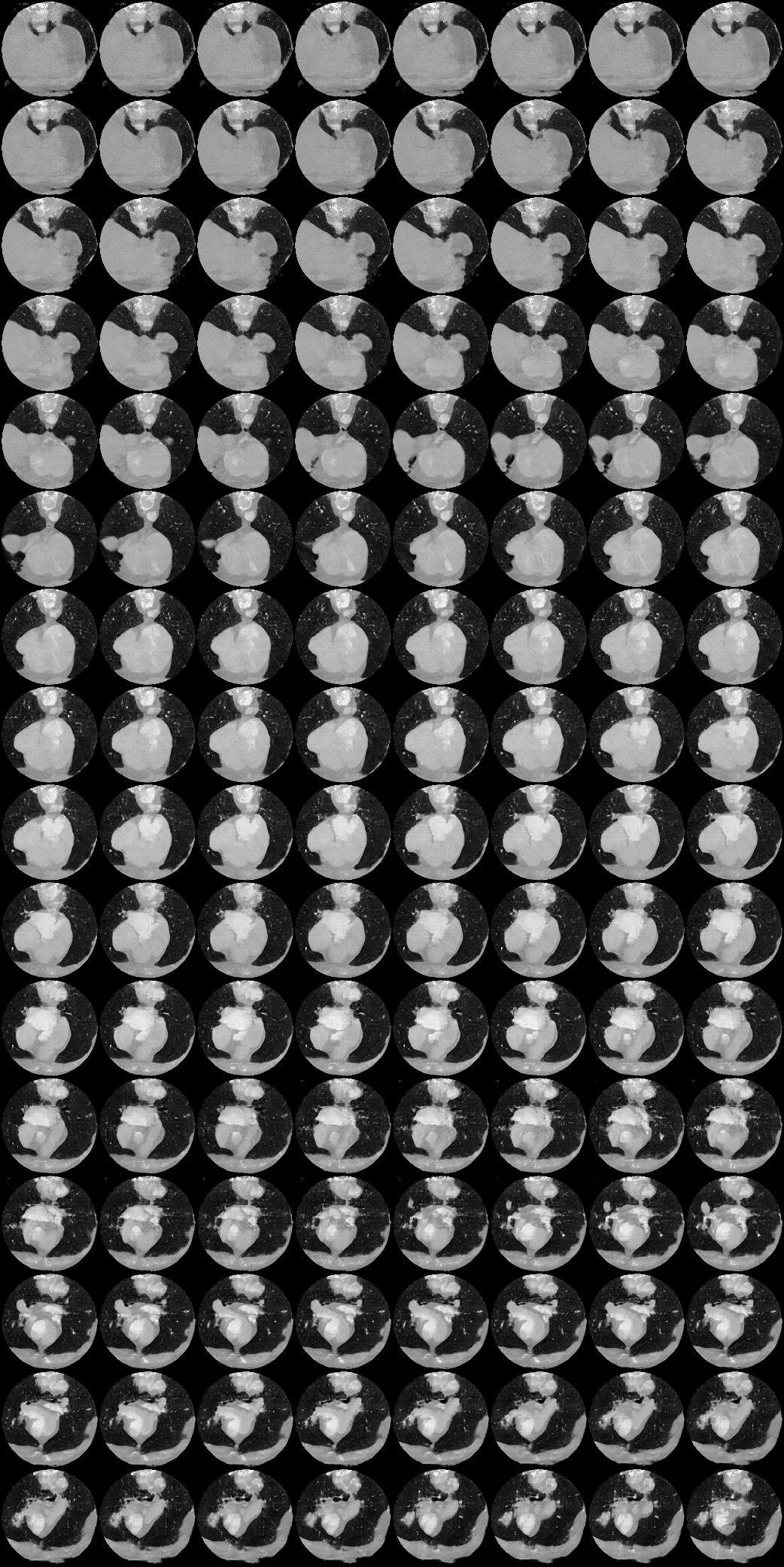
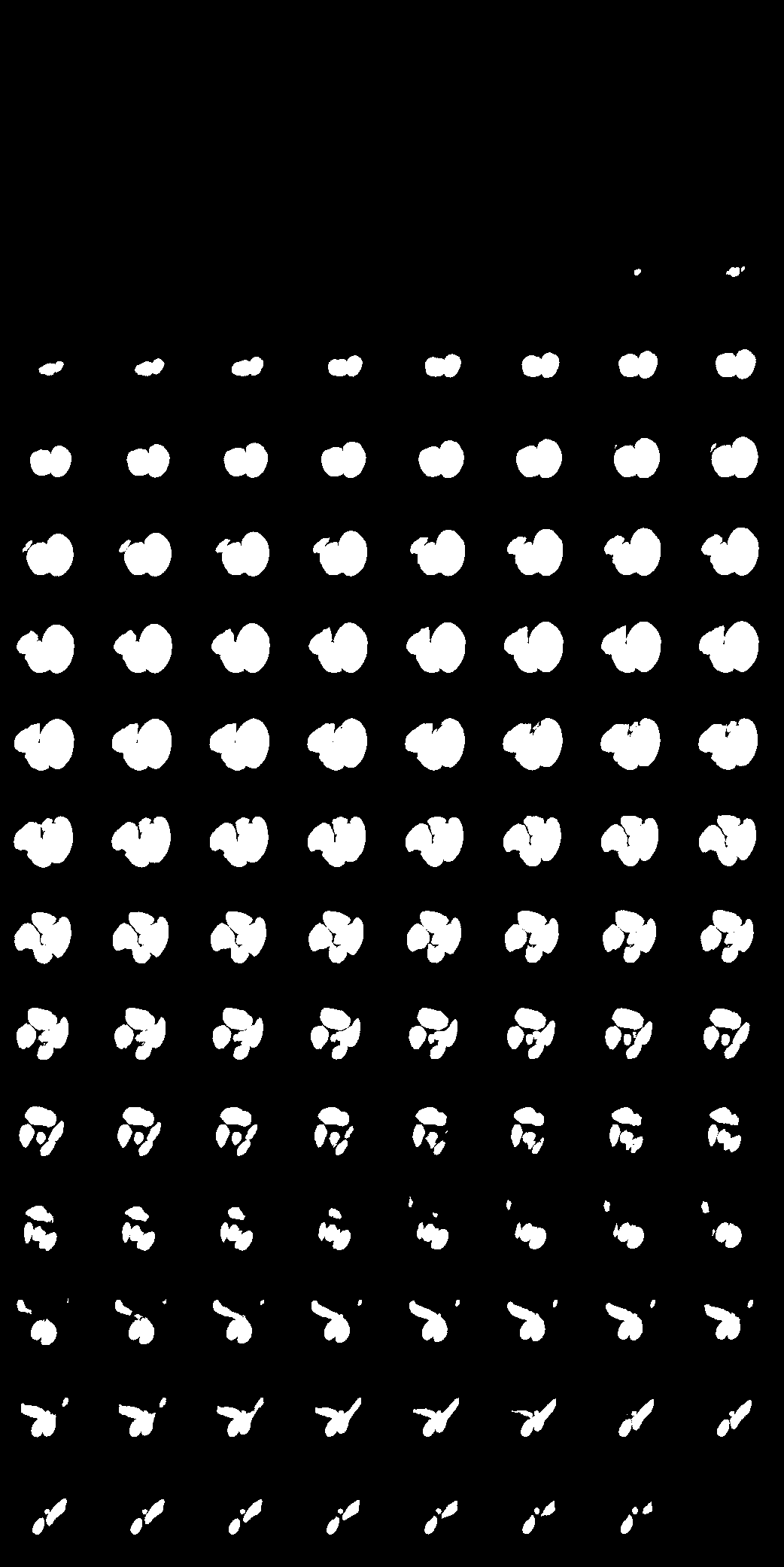
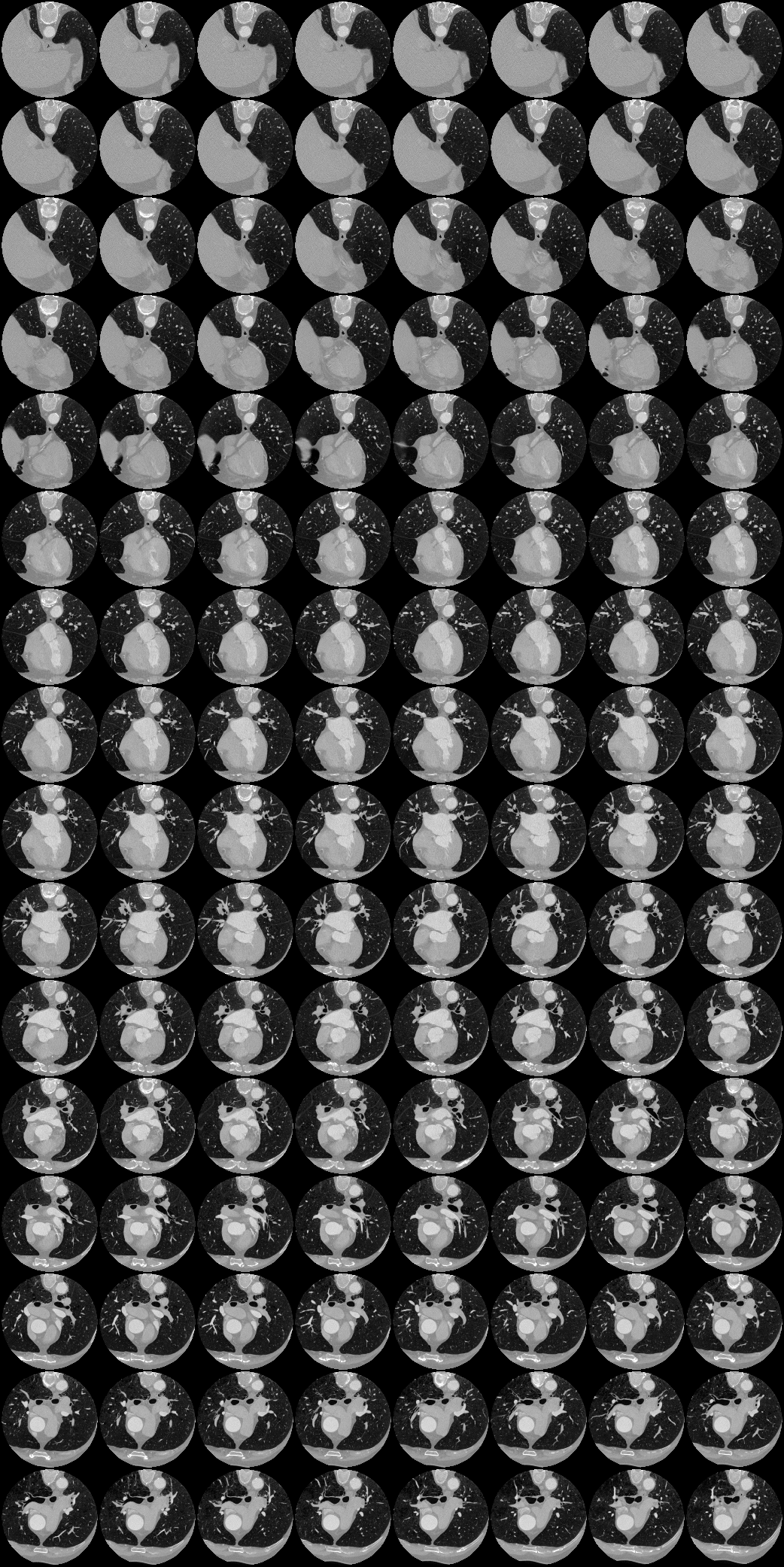
Labelling data is expensive and time consuming especially for domains such as medical imaging that contain volumetric imaging data and require expert knowledge. Exploiting a larger pool of labeled data available across multiple centers, such as in federated learning, has also seen limited success since current deep learning approaches do not generalize well to images acquired with scanners from different manufacturers. We aim to address these problems in a common, learning-based image simulation framework which we refer to as Federated Simulation. We introduce a physics-driven generative approach that consists of two learnable neural modules: 1) a module that synthesizes 3D cardiac shapes along with their materials, and 2) a CT simulator that renders these into realistic 3D CT Volumes, with annotations. Since the model of geometry and material is disentangled from the imaging sensor, it can effectively be trained across multiple medical centers. We show that our data synthesis framework improves the downstream segmentation performance on several datasets.
 |
Daiqing Li, Amlan Kar, Nishant Ravikumar, Alejandro F Frangi, Sanja Fidler
Federated Simulation for Medical Imaging MICCAI, 2020. (Early Accept) |
|
|
|
|
|
|
|
|
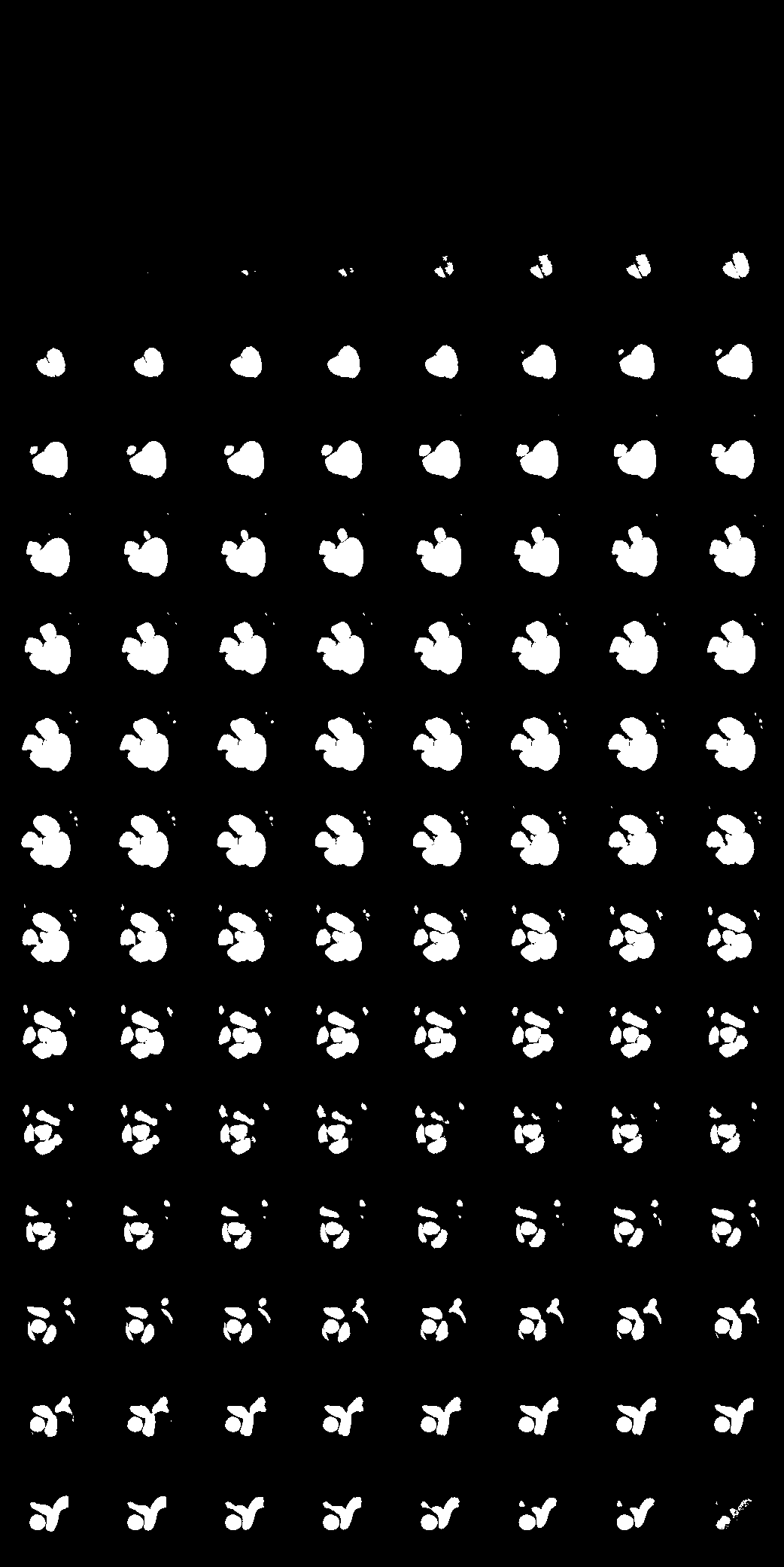
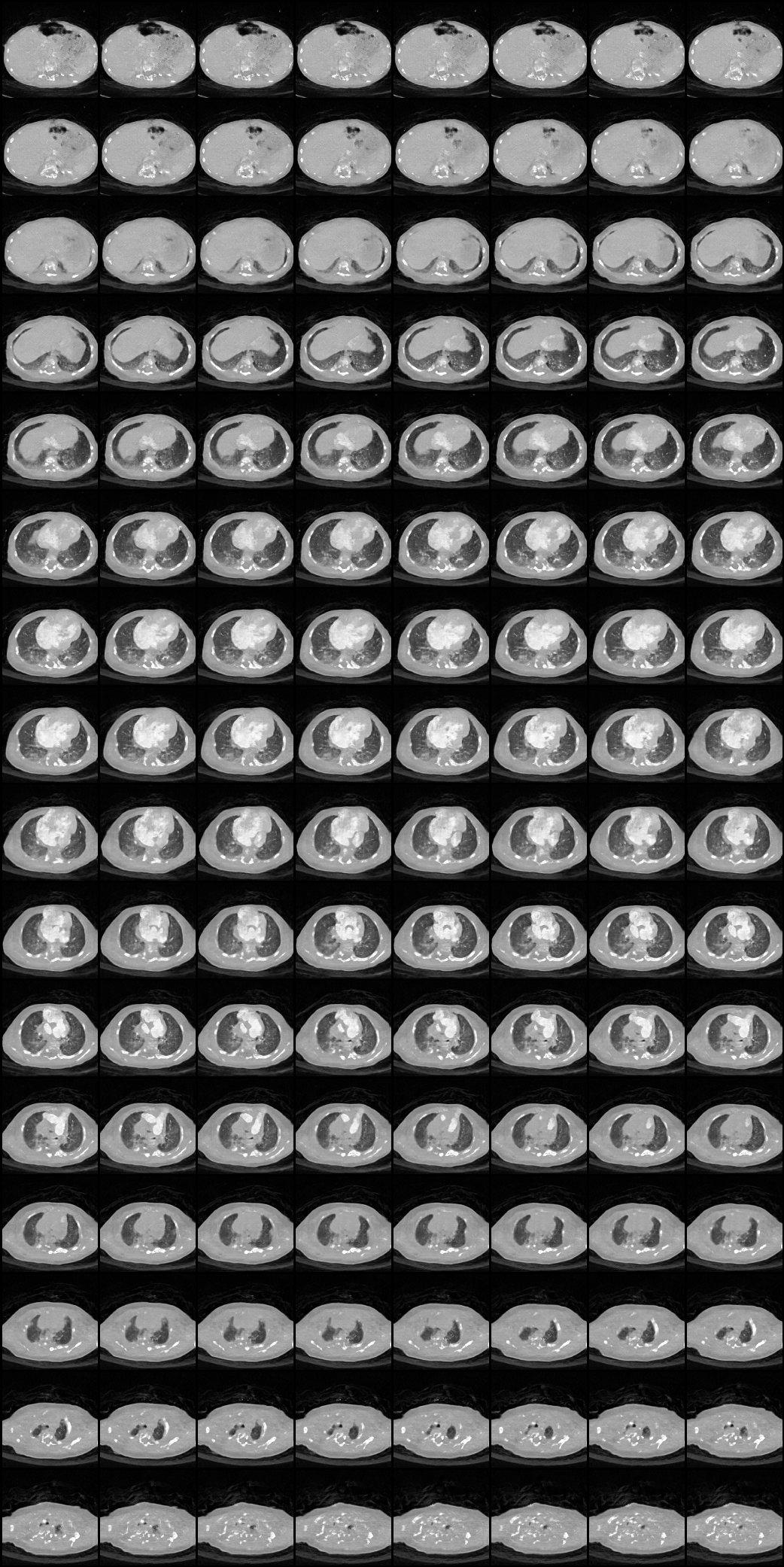
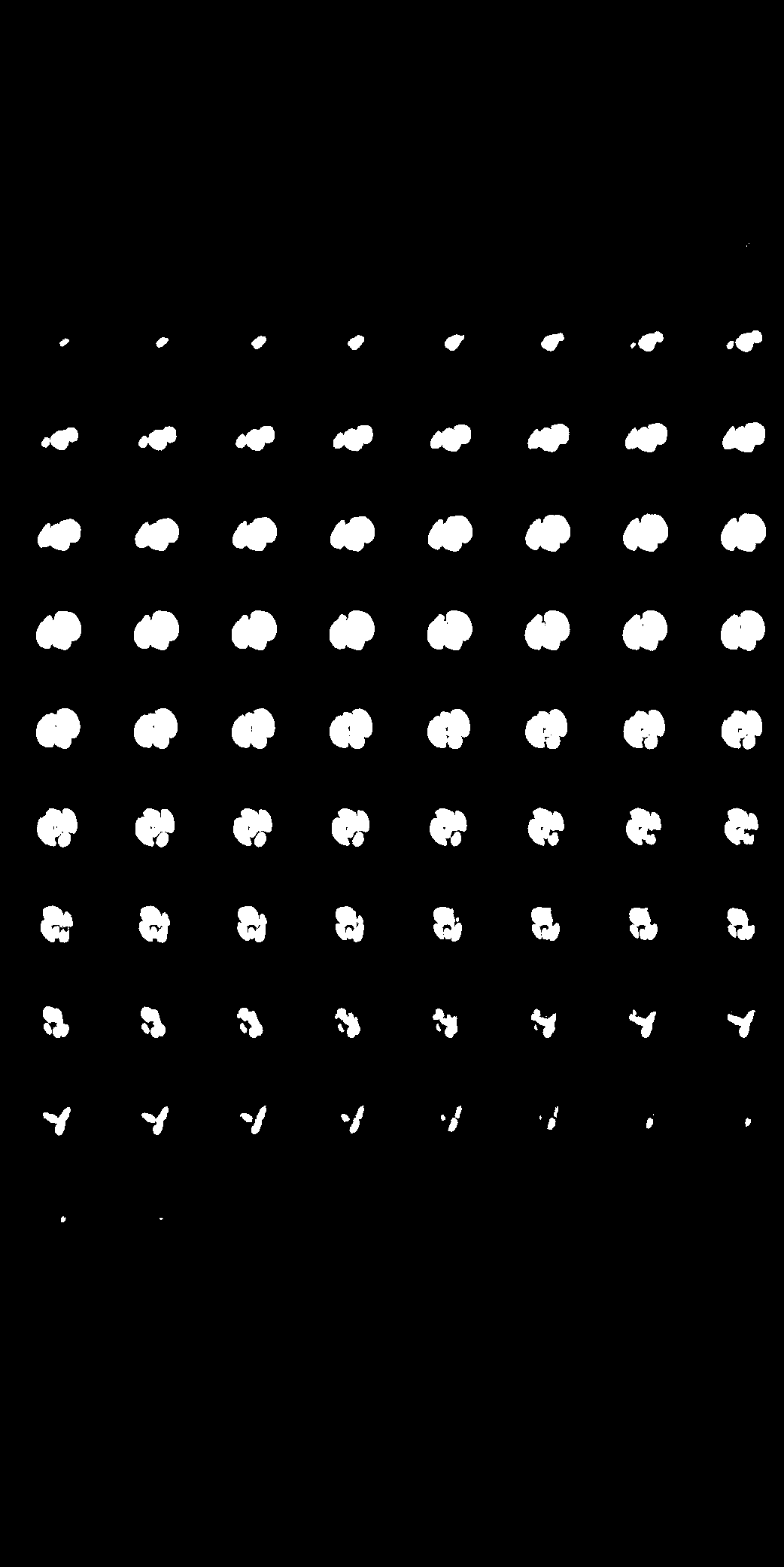
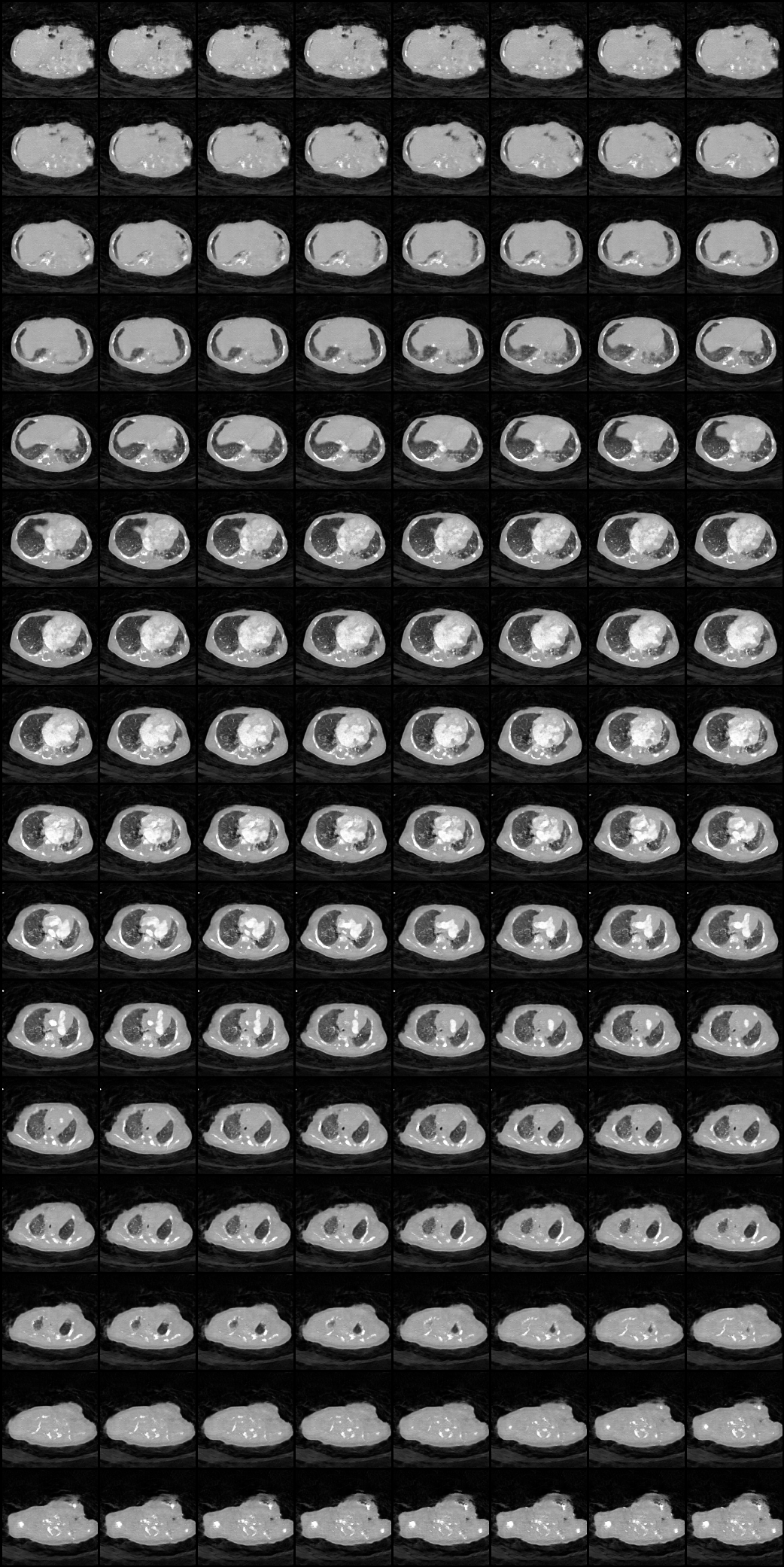
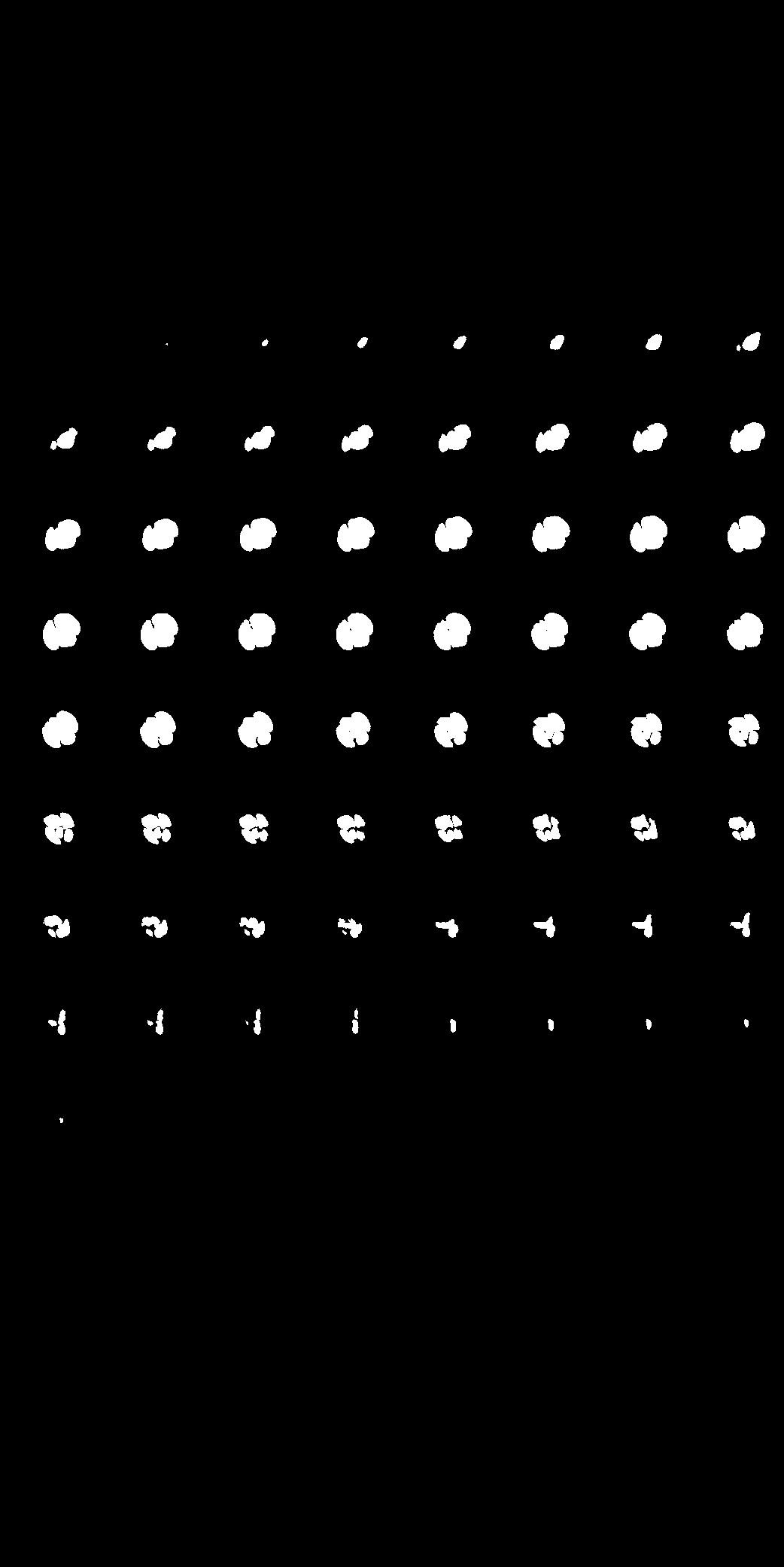
| Here we showcase an interactive simulation application of our approach using NVIDIA Omniverse Kit. The users can adjust the shape parameters of a cardiac mesh template and synthesize a novel (labeled) cardiac CT volume. Users can view different slices along the z axis by adjusting the slice bar. Notice that the generated volume and the input mesh are consistent across slices. |
|
|
|
|
| Presentation video at MICCAI 2020. |
| Medical data usually lives at multiple sites, and sharing is a problem due to patient privacy. In our work, we learn to synthesize labeled CT volumes in a federated setting that is privacy preserving. In the FL setup, we assume there is a central server that aggregates and maintains a global model for the organ shape and material across sites, while each individual site has a local simulator with specific device parameters and its own conditional GAN. The central server shares the global knowledge of shape and material parameters across sites, which they use to train their enhancer using local data. This setting only requires transmitting gradients with respect to the shape and material model parameters to learn a global simulator, which we hypothesize should contain minimal information about individual patients. To train models for downstream tasks, every site can simulate their own data and train their own unique task model, as opposed to traditional FL where one model architecture and weights must be used and shared across sites and gradients for the model itself must be transmitted. |
 |
 |
 |
 |
||||
 |
 |
 |
 |
||||
 |
 |
 |
 |